Compare commits
10 Commits
| Author | SHA1 | Date | |
|---|---|---|---|
| 295d244aaf | |||
| 55f3d7a80d | |||
| c025e87fbb | |||
| d480fe92b7 | |||
| ca7be61d56 | |||
| 177f63e774 | |||
| 9e2ad9c7af | |||
| c6e80fc71a | |||
|
|
95cf42c513 | ||
| 1223366400 |
91
README.md
91
README.md
@ -1,28 +1,64 @@
|
||||
# Erlang Decision Tree and Baysian Networks
|
||||
|
||||

|
||||
|
||||
This repository have two different exercises in erlang:
|
||||
|
||||
**1.Decision tree that maximizes the optimal point and consequently provides decision support based on the assumptions provided.**
|
||||
**1.Decision tree that maximizes the optimal point and consequently provides decision support based on the given assumptions.**
|
||||
|
||||
* The probability of making a medication/treatment 1 and use is OK of 42%;
|
||||
* One possibility of making a medication/treatment 1 and of the user staying KO is 22.5%;
|
||||
* The probability of making a medication/treatment 1 and is not conclusive is 35.6%;
|
||||
* The probability of making a medication/treatment 2 and staying OK is 62%;
|
||||
* The probability of making a medication/treatment 2 and getting KO is 38%.
|
||||
* The probability of taking medication/treatment 1 and the user being OK is 42%;
|
||||
* The probability of taking medication/treatment 1 and the user becoming KO is 22.5%;
|
||||
* The probability of taking medication/treatment 1 and it not being conclusive is 35.6%;
|
||||
* The probability of taking medication/treatment 2 and being OK is 62%;
|
||||
* The probability of taking a medication/treatment 2 and getting KO is 38%.
|
||||
|
||||

|
||||
```
|
||||
|
||||
-module(engine).
|
||||
-compile([export_all]).
|
||||
|
||||
% Decision Tree
|
||||
%
|
||||
% +----+
|
||||
% +---> OK |
|
||||
% | +----+
|
||||
% |
|
||||
% +----------| +----+
|
||||
% | DO MED 1 +---> KO |
|
||||
% +----------+ +----+ +----+
|
||||
% | +---> OK |
|
||||
% | +---------------| +----+
|
||||
% +---> DO MED 2 +
|
||||
% +---------------| +----+
|
||||
% +---> KO |
|
||||
% +----+
|
||||
% Each edge can be calculated from the table
|
||||
|
||||
|
||||
doAll() ->
|
||||
T = dec_tree:start(),
|
||||
T ! {create_node, 'DO MED1', 0, none},
|
||||
T ! {create_node, 'MED1 OK', 0.980, 'DO MED1'},
|
||||
T ! {create_node, 'MED1 KO', 0.225, 'DO MED1'},
|
||||
T ! {create_node, 'DO MED2', 0.356, 'DO MED1'},
|
||||
T ! {create_node, 'MED 2 OK', 0.62, 'DO MED2'},
|
||||
T ! {create_node, 'MED 2 KO', 0.38, 'DO MED2'},
|
||||
T ! {solve}.
|
||||
|
||||
|
||||
```
|
||||
|
||||
|
||||
|
||||
|
||||
|
||||
**2.There are ways to get the probabilities knowing certain parameters (they do not indicate the optimal point, but calculate a probability of success depending on the existing resources) - using the baysian networks.**
|
||||
**2. There are ways of obtaining probabilities knowing certain parameters (they don't indicate the optimum point, but calculate a probability of success depending on the existing resources) - using Baysian networks.**
|
||||
|
||||
What is the probability of giving the patient M2 and is OK, knowing that:
|
||||
What is the probability of giving the patient M2 and the patient being well, knowing that:
|
||||
|
||||
* The probability of administering M1 and the patient is OK is 20%;
|
||||
* The probability of administering M2 knowing that I administered M1 and the patient being OK is 70%;
|
||||
* The probability of administering M2 knowing that I did not administer M1 and the patient is OK of 20%.
|
||||
* The probability of administering M1 and the patient being well is 20 per cent;
|
||||
* The probability of administering M2 knowing that I have administered M1 and the patient is well is 70%;
|
||||
* The probability of administering M2 knowing that I have not administered M1 and the patient is OK is 20 per cent.
|
||||
|
||||
### Prerequisites
|
||||
|
||||
@ -33,12 +69,12 @@ Basic knowledge of erlang, algorithm and statistics.
|
||||
|
||||
### Install or use Docker
|
||||
|
||||
If you prefer install the erlang compiler, please search for the appropriate installation for your OS.
|
||||
If you prefer to install the erlang compiler, please find the appropriate installation for your operating system.
|
||||
|
||||
If you prefer you can use Docker to test this solution.
|
||||
Get a [docker container](https://hub.docker.com/search?q=erlang&type=image) and start the container:
|
||||
I suggest [this container](https://hub.docker.com/r/bitwalker/alpine-erlang) and run:
|
||||
If you prefer, you can use Docker to test this solution.
|
||||
Get a [docker container](https://hub.docker.com/search?q=erlang&type=image) and start the container.
|
||||
|
||||
I suggest [this container](https://hub.docker.com/r/bitwalker/alpine-erlang) and run it:
|
||||
|
||||
```
|
||||
docker pull bitwalker/alpine-erlang
|
||||
@ -50,15 +86,15 @@ docker run --rm -it --user=root bitwalker/alpine-erlang
|
||||
1. For the first exercise (Decision Tree):
|
||||
|
||||
```
|
||||
c(engine).
|
||||
engine:doAll().
|
||||
c(motor).
|
||||
motor:doAll().
|
||||
```
|
||||
|
||||
and the best solution is:
|
||||
|
||||
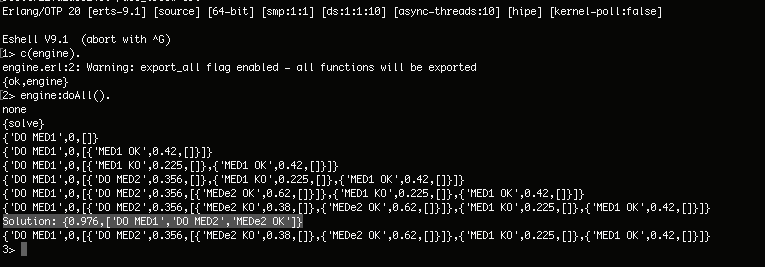
|
||||
|
||||
** You can do a medication 1 and medication 2 of 97.6% of success to be OK.**
|
||||
** You can take medication 1 and medication 2 with a 97.6 per cent success rate to get well.
|
||||
|
||||
|
||||
|
||||
@ -67,16 +103,16 @@ and the best solution is:
|
||||
2. For the second exercise (Baysian Networks):
|
||||
|
||||
```
|
||||
c(engine3).
|
||||
engine3:getp({m2, ok}).
|
||||
c(motor3).
|
||||
motor3:getp({m2, ok}).
|
||||
```
|
||||
|
||||
|
||||
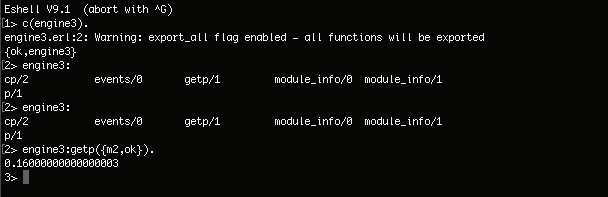
|
||||
|
||||
**There is a 16% chance that the patient will be OK administering the drug M2.**
|
||||
**There is a 16 per cent chance that the patient will be OK when administering drug M2.
|
||||
|
||||
## Built With
|
||||
## Created with
|
||||
|
||||
* erlang - https://www.erlang.org;
|
||||
|
||||
@ -86,13 +122,12 @@ Please read [CONTRIBUTING.md](CONTRIBUTING.md) for details on my code of conduct
|
||||
|
||||
## Versioning
|
||||
|
||||
I use [SemVer](http://semver.org/) for versioning. For the versions available, see the [tags on this repository](https://gitlab.andrealmeida.net/public_projects/erlang-decision-tree/tags).
|
||||
I use [SemVer](http://semver.org/) for versioning. For available versions, see [tags on this repository](https://gitlab.andrealmeida.net/public_projects/erlang-decision-tree/tags).
|
||||
|
||||
## Authors
|
||||
|
||||
* **André Almeida** - [andrealmeida.net](https://andrealmeida.net)
|
||||
[](https://andrealmeida.net)
|
||||
|
||||
## License
|
||||
|
||||
This project is licensed under the MIT License - see the [LICENSE.md](LICENSE.md) file for details
|
||||
## Licence
|
||||
|
||||
This project is licensed under the MIT licence - see the file [LICENSE.md](LICENSE.md) for more details.
|
||||
|
||||
29
app/baysian-network/engine3.erl
Normal file
29
app/baysian-network/engine3.erl
Normal file
@ -0,0 +1,29 @@
|
||||
-module(engine3).
|
||||
-compile([export_all]).
|
||||
|
||||
events() -> [{m1, ok}, {m2, ok}].
|
||||
|
||||
p({m1, ok}) -> 0.2;
|
||||
p(A) when is_tuple(A) -> none.
|
||||
|
||||
cp({m2, ok}, {m1, ok}) -> 0.8;
|
||||
cp(_, _) -> none.
|
||||
|
||||
getp({A, B}) when is_tuple(A), is_tuple(B) ->
|
||||
case cp(A, B) of
|
||||
none ->
|
||||
Pba = cp(B, A),
|
||||
Pa = getp(A),
|
||||
Pb = getp(B),
|
||||
Pba * Pa / Pb;
|
||||
X ->
|
||||
X
|
||||
end;
|
||||
|
||||
getp(A) when is_tuple(A) ->
|
||||
case p(A) of
|
||||
none ->
|
||||
lists:sum([ getp({A, B})*getp(B) || B <- events(), p(B) /= none, cp(A,B) /= none ]);
|
||||
X ->
|
||||
X
|
||||
end.
|
||||
41
app/decision-tree/dec_tree.erl
Normal file
41
app/decision-tree/dec_tree.erl
Normal file
@ -0,0 +1,41 @@
|
||||
-module(dec_tree).
|
||||
-export([start/0, decision_tree/0]).
|
||||
|
||||
%%% Interface
|
||||
|
||||
start() -> Pid = spawn(?MODULE, decision_tree, []), Pid.
|
||||
|
||||
decision_tree() -> T = create_tree(), decision_tree(T).
|
||||
decision_tree(Tree) ->
|
||||
io:format("~w~n", [Tree]), %% for debug purposes we always print the Tree
|
||||
receive
|
||||
{create_node, Node, Edge, Parent} -> NewTree = create_node(Tree, Node, Edge, Parent), decision_tree(NewTree);
|
||||
{print} -> decision_tree(Tree); % no need to print the tree, since we already printed before receive...
|
||||
{solve} -> io:format("Solution: ~w~n", [solve(Tree)]), decision_tree(Tree)
|
||||
end.
|
||||
|
||||
%%% Create the decision Tree
|
||||
|
||||
create_tree() -> none.
|
||||
|
||||
create_node(none, Node, Edge, _) -> {Node, Edge, []}; % 1st node
|
||||
create_node({Parent, P_Edge, L}, Node, Edge, Parent) -> {Parent, P_Edge, [{Node, Edge, []}|L]}; % we find the parent node and insert the new node
|
||||
create_node({Root, R_Edge, []}, _, _, Parent) -> {Root, R_Edge, []}; % recursion stop when the parent node is not found
|
||||
create_node({Root, R_Edge, L}, Node, Edge, Parent) -> {Root, R_Edge, lists:map(fun(N) -> create_node(N, Node, Edge, Parent) end, L)}. % we try to insert the node in all subtrees
|
||||
|
||||
%%% Solve the decision Tree
|
||||
|
||||
% Determines the path by choosing the greatest edge
|
||||
solve_max([{Edge, Path}]) -> {Edge, Path};
|
||||
solve_max([{H_Edge, E_Path}|T]) ->
|
||||
{T_max, T_Path} = solve_max(T),
|
||||
if
|
||||
H_Edge > T_max ->
|
||||
{H_Edge, E_Path};
|
||||
true -> % works as an 'else' branch
|
||||
{T_max, T_Path}
|
||||
end.
|
||||
|
||||
solve(none) -> false; %there is no solition for an empty decision tree
|
||||
solve({Node, Edge, []}) -> {Edge,[Node]}; %recursion stop
|
||||
solve({Node, Edge, L}) -> {C_Edge, C_Path} = solve_max(lists:map(fun(N) -> solve(N) end, L)), {Edge + C_Edge, [Node|C_Path]}.
|
||||
30
app/decision-tree/engine.erl
Normal file
30
app/decision-tree/engine.erl
Normal file
@ -0,0 +1,30 @@
|
||||
-module(engine).
|
||||
-compile([export_all]).
|
||||
|
||||
% Decision Tree
|
||||
%
|
||||
% +----+
|
||||
% +---> OK |
|
||||
% | +----+
|
||||
% |
|
||||
% +----------| +----+
|
||||
% | DO MED 1 +---> KO |
|
||||
% +----------+ +----+ +----+
|
||||
% | +---> OK |
|
||||
% | +---------------| +----+
|
||||
% +---> DO MED 2 +
|
||||
% +---------------| +----+
|
||||
% +---> KO |
|
||||
% +----+
|
||||
% Each edge can be calculated from the table
|
||||
|
||||
|
||||
doAll() ->
|
||||
T = dec_tree:start(),
|
||||
T ! {create_node, 'DO MED1', 0, none},
|
||||
T ! {create_node, 'MED1 OK', 0.980, 'DO MED1'},
|
||||
T ! {create_node, 'MED1 KO', 0.225, 'DO MED1'},
|
||||
T ! {create_node, 'DO MED2', 0.356, 'DO MED1'},
|
||||
T ! {create_node, 'MED 2 OK', 0.62, 'DO MED2'},
|
||||
T ! {create_node, 'MED 2 KO', 0.38, 'DO MED2'},
|
||||
T ! {solve}.
|
||||
Loading…
Reference in New Issue
Block a user